Further reading :
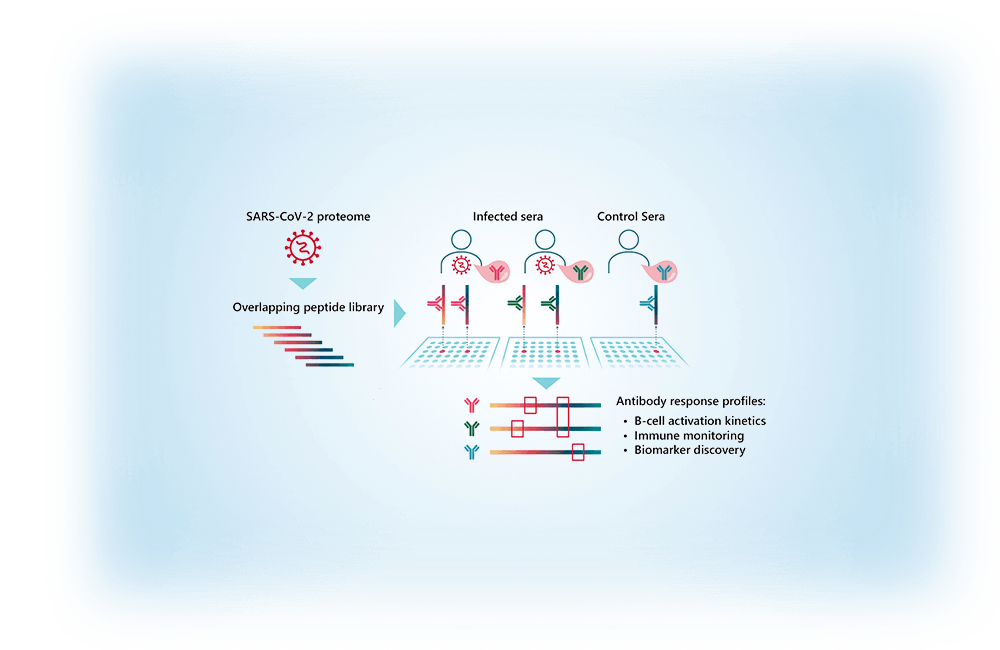
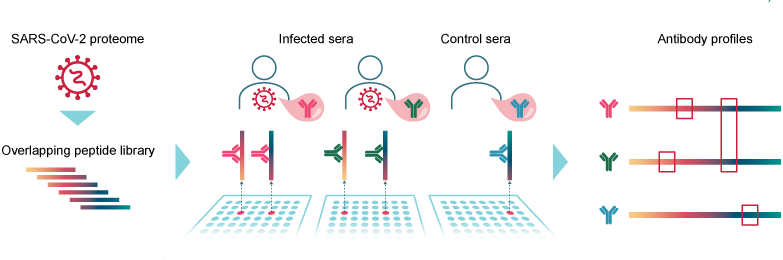
SARS-CoV-2 proteome-wide analysis revealed significant epitope signatures in COVID-19 patients.
Tatjana Schwarz, Kirsten Heiss, Yuvaraj Mahendran, Fiordiligie Casilag, Florian Kurth, Leif Sander, Clemens-Martin Wendtner, Manuela A. Hoechstetter, Marcel A. Müller, Renate Sekul, Christian Drosten, Volker Stadler and Victor M. Corman. Front. Immunol. [https://www.frontiersin.org/articles/10.3389/fimmu.2021.629185/abstract]
SARS-CoV-2 Spike Protein Arrested in the Closed State Induces Potent Neutralizing Responses.
George W. Carnell, Katarzyna A. Ciazynska, David A. Wells, Xiaoli Xiong, Ernest T. Aguinam, Stephen H. McLaughlin, Donna Mallery. Preprint. Immunology, 14. January 2021. [doi.org/10.1101/2021.01.14.426695]
SARS-CoV-2 Epitope Mapping on Microarrays Highlights Strong Immune-Response to N Protein Region.
Angelo Musicò, Roberto Frigerio, Alessandro Mussida, Luisa Barzon, Alessandro Sinigaglia, Silvia Riccetti, Federico Gobbi. Vaccines 9, Nr. 1 (11. January 2021): 35. [doi.org/10.3390/vaccines9010035]
Recent Advances in the Diagnosis of COVID-19: A Bird’s Eye View.
Bhawna Sharma, Mohd FardeenHusain Shahanshah, Sanjay Gupta, and Vandana Gupta. Expert Review of Molecular Diagnostics, 10. January 2021, 14737159.2021.1874354. [doi.org/10.1080/14737159.2021.1874354]
Generation of Chicken IgY against SARS-COV-2 Spike Protein and Epitope Mapping.
Yan Lu, Yajun Wang, Zhen Zhang, Jingliang Huang, Meicun Yao, Guobin Huang, Yuanyuan Ge, et al. Published by Juraj Ivanyi. Journal of Immunology Research 2020 (16. October 2020): 1–8. [doi.org/10.1155/2020/9465398]
Rapid Response to Pandemic Threats: Immunogenic Epitope Detection of Pandemic Pathogens for Diagnostics and Vaccine Development Using Peptide Microarrays.
Kirsten Heiss, Jasmin Heidepriem, Nico Fischer, Laura K Weber, Christine Dahlke, Thomas Jaenisch, und Felix F Loeffler. Journal of Proteome Research, 5. September 2020 [doi.org/10.1021/acs.jproteome.0c00484]
Molecular and Serological Tests for COVID-19. A Comparative Review of SARS-CoV-2 Coronavirus Laboratory and Point-of-Care Diagnostics.
Kubina, Robert, und Arkadiusz Dziedzic. Diagnostics 10, Nr. 6 (26. June 2020): 434. [doi.org/10.3390/diagnostics10060434]
Distinct Early IgA Profile May Determine Severity of COVID-19 Symptoms: An Immunological Case Series.
Dahlke, Christine, Jasmin Heidepriem, Robin Kobbe, Rene Santer, Till Koch, Anahita Fathi, My L. Ly, et al. Preprint. Infectious Diseases (except HIV/AIDS), April 17, 2020 [doi.org/10.1101/2020.04.14.20059733]
IgG and IgA Antibody Profiling with the PEPperCHIP® Infectious Disease Epitope Microarray. (Application note)
Discovery of serotype-specific epitope biomarkers for the diagnosis of Dengue virus infections. (Poster)
Profiling epitope-specific antibody responses against malaria utilizing high-density peptide arrays. (Poster)