Join Us For Our 9th ibiTea - A Scientific Get Together

Are you interested in learning about…
A Microscopy-Based CRISPR Screening Platform?
If so, grab a cup of tea and join our scientific discussion on:
15 November 2023, 10PM SGT
The number of participants is limited, so please reserve your seat now.
ibiTea Topic Outline
SPARCS, a Platform for Genome-Scale CRISPR Screening for Subcellular Spatial Phenotypes
Presented by Niklas Schmacke, Gene Center, LMU Munich, Germany (2023 ibidi Paper Award Winner)
In our 9th ibiTea, Niklas Schmacke will present a new microscopy-based CRISPR screening platform:
Forward genetic screening associates phenotypes with genotypes by randomly inducing mutations and then identifying those that result in phenotypic changes of interest. Niklas and his team have developed “spatially resolved CRISPR screening” (SPARCS), a platform for microscopy-based genetic screening for spatial cellular phenotypes.
SPARCS uses automated high-speed laser microdissection to physically isolate phenotypic variants in situ from virtually unlimited library sizes. They demonstrate the potential of SPARCS in a genome-wide CRISPR-KO screen on autophagosome formation in 40 million cells.
Coupled with deep learning image analysis, SPARCS recovered almost all known macroautophagy genes in a single experiment and discovered a role for the ER-resident protein EI24 in autophagosome biogenesis. Harnessing the full power of advanced imaging technologies, SPARCS enables genome-wide forward genetic screening for diverse spatial phenotypes in situ.
Read the preprint of the full article here.
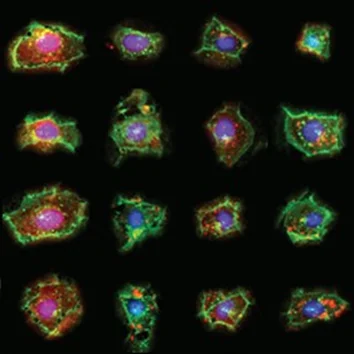
A collage of single U2OS cells with different genotypes treated with
Torin-1 or left untreated.
Red: mCherry-LC3. Green: Cell membranes stained with wheat germ agglutinin. Blue: Nuclei.
Speaker
Niklas Schmacke
Gene Center, LMU Munich, Germany
Niklas Schmacke, during his PhD in Veit Hornung’s lab, discovered a new priming modality of the NLRP3 inflammasome, a cell death and inflammation inducing protein complex that plays a critical role in diverse pro-inflammatory contexts, including infection and atherosclerosis. He also developed SPARCS, a high-resolution genome-wide CRISPR screening technology for image-based phenotypes. Currently, he works with Fabian Theis and Veit Hornung on the unbiased identification of subcellular spatial phenotypes in single cell images using deep learning.
